Implementing distributed grid search for deep learning using TensorFlow, scikit-learn, and joblib¶
What is this talk about?¶
It's about combining open source technologies to implement a machine learning workflow.
- custom machine learning
- TensorFlow
- a standard interface for machine learning
- scikit-learn
- simple distributed computing
- joblib
1. Custom machine learning¶
- ML implementation is getting easier, almost like writing equations on a whiteboard.
- The focus here: supervised, discriminative deep learning with gradient-based optimization
- alternatives: tree ensembles, probabilistic graphical models (PyMC3, STAN), etc.
Procedure for finding a good $f$¶
Typically, we use gradient-based optimization.
- Define an objective function $g(X, y, W)$
- $X$, $y$: training data (fixed)
- Goal: find setting for $W$ that optimizes $g$.
- Often, $g$ is a difference between $\hat{y} = f(y \mid X, W)$ and $y$.
- Define a function to compute the derivative of $g$ with respect to $W$.
- This often duplicates engineering effort from implementing $f$ and $g$.
- For DL models, $g$ and its derivative can be very complicated.
- Iterate:
- Compute the derivative.
- Update $W$ based on the derivative.
- Often with complex adjustments to reduce required iterations.
Graph-based numerical computation (e.g., TensorFlow)¶
- step 1 (model and objective implementation) becomes pretty easy
- step 2 (differentiation) becomes one line of code
- step 3 (iterative learning) becomes one or a few lines of code
TensorFlow and the alternatives¶
- TensorFlow's graph-based approach to numerical computation facilitates DL implementation.
- Developed by Google, now an open source project.
- It's pretty popular:
There are alternative DL tools, in Python (often with C++ underneath) and other languages.
- theano (Python)
- very similar predecessor to TF
- academic research project
- keras (Python)
- wraps TF and theano
- mainly for neural nets & deep learning
- mxnet (Python)
- torch (Lua)
- just NumPy + SciPy
scipy.optimizeis handy for many things- no autodiff
- caffe
- pycaffe
- Microsoft CNTK
- others
- there's probably a new one as you read this slide.
A brief introduction to TensorFlow¶
In [22]:
"""
This is an example of accumulating a sum of 1-D tensors,
to illustrate some major concepts in TensorFlow
(variables, operations, graphs, and sessions).
"""
import numpy as np
import tensorflow as tf
# A graph is a collection of tensors and operations on them.
g = tf.Graph()
# When you create tensors and operations, they are added to
# whatever the current default graph is.
# TF lets you use context managers rather than, e.g.,
# having to do something like g.add(variable).
# Note: there is also a global default graph (global scope).
with g.as_default():
# Define a placeholder for input (which gets added to g).
x = tf.placeholder(dtype=np.float32, name="x")
# Make a variable, including an initializer (here just a constant value).
# Its value will be kept across calls to `run` within a session (more below).
sum_x = tf.Variable([0., 0., 0.], name="sum_x")
# You need to make sure variables are initialize in a session.
init_op = tf.initialize_all_variables()
# Make an Operation to add the input to the sum.
# This is also added to the graph.
# Note: the value returned by this op this will be `sum_x`.
add_x = sum_x.assign(sum_x + x)
# Sessions are execution contexts. They capture, e.g.,
# the values of Variable instances in the graph.
# Analogy: think of a tf.Graph as like a UNIX script/program/executable
# and a tf.Session as like a UNIX process. It's not a perfect analogy.
with tf.Session() as sess:
# Sessions consist of multiple runs, so perhaps the run sequence
# is the script in the analogy above.
sess.run(init_op)
print("session 1, initial:",
sess.run(sum_x))
print("session 1, after call 1:",
sess.run(add_x, feed_dict={x: [1, 2, 3]}))
print("session 1, after call 2:",
sess.run(add_x, feed_dict={x: [0, 5, 0]}))
print("session 1, final:",
sess.run(add_x, feed_dict={x: [0, -10, 0]}))
session 1, initial: [ 0. 0. 0.] session 1, after call 1: [ 1. 2. 3.] session 1, after call 2: [ 1. 7. 3.] session 1, final: [ 1. -3. 3.]
In [23]:
# Now start another session. The previous state (variable values) is lost.
# Note how we don't have to double indent as above.
with g.as_default(), tf.Session() as sess:
# If `init_op` isn't called again, we get an exception because
# the variable hasn't been initialized.
sess.run(init_op)
print("session 2, initial sum_x:", sess.run(sum_x))
print("session 2, after call 1:",
sess.run(add_x, feed_dict={x: [1, 1, 1]}))
session 2, initial sum_x: [ 0. 0. 0.] session 2, after call 1: [ 1. 1. 1.]
In [24]:
"""
Here's a quick illustration of automatic differentiation.
Note that TensorFlow implements learning algorithms that use this internally,
to make it easy to apply various gradient-based optimizers.
"""
g = tf.Graph()
with g.as_default(), tf.Session() as sess:
a = tf.constant([[2.0, 5.0]])
x = tf.Variable([[0.0, 0.0]])
y = a * x + 10.0
y_grad = tf.gradients([y], [x])[0] # inputs/outputs lists
sess.run(tf.initialize_all_variables())
print("Derivative of y = a * x + 10.0 with respect to x:\n",
sess.run(y_grad))
Derivative of y = a * x + 10.0 with respect to x: [[ 2. 5.]]
Multilayer Perceptrons (MLPs)¶
- a feed-forward neural network
- e.g., no recursion, no convolutions
- input layer, 1+ hidden layers, output layer
- can model interactions between features
- capable of approximating any function ("universal approximation theorem")
- an old idea (Rosenblatt, 1961), but can be used with state-of-the-art methods
- e.g., ReLUs, dropout, Adam learning algorithm
Notes:
- Here, we have a softmax output function $f$, for multiclass classification.
- ReLU hidden units could also be used, but they might be less interpretable than sigmoid hidden units.
2. A standard interface for machine learning¶
We want to follow a standard to facilitate:
- use in existing scikit-learn-based workflows
- easy comparisons to other methods in scikit-learn
The scikit-learn Estimator API¶
- scikit-learn is a ML library developed by many people
- The consistent API makes trying different ML algorithms easy.
- It allows users to implement their own estimators.
- helper functions for input validation, preprocessing, testing, etc.
- key feature:
sklearn.utils.estimator_checks.check_estimatorto check API conformity of a custom estimator (free tests!)
- It facilitates 2 big ML challenges:
- pipelines of transformations
- e.g., extract features, transform/standardize values, fit model
- gridsearch over hyperparameters (more on this later)
- pipelines of transformations
Overview for custom scikit-learn predictive models¶
- For models, we need to implement a
fit(X, y)andpredict(X)- optionally, also
predict_proba(X), etc.
- optionally, also
__init__should just attach argumentsfitdoes a lot of what__init__normally doesfitusually sets instance attributes (e.g.,model.coef_)
- everything should be serializable with
picklejoblib, used by grid search, expects things to be serializable
- mixins for different model types
RegressorMixin,ClassifierMixin
The Civis MLP Implementation¶
- follows scikit-learn's API
- GPU support through TensorFlow
- sparse and dense matrix support
- generic base class + regressor and classifier subclasses
- classifier supports binary, multiclass, and multilabel binary classification
- called MuFFNN (Multilayer Feed-Forward Neural Network)
A few alternative MLP implementations¶
sklearn.neural_network.MLPClassifier(andMLPRegressor)- CPU-only, no GPU support
- may be difficult to customize (e.g., new learning algorithms)
tensorflow.learn- included with TensorFlow in a
contribsection - doesn't quite follow scikit-learn interface (last I checked)
- e.g.,
__init__does more than attach arguments - possibly difficult to use with grid search and pipelines
- e.g.,
- extra support for out-of-core, minibatch learning (
DataFeederclasses)
- included with TensorFlow in a
keras.wrappers.scikit_learn.KerasClassifier- If you implement a model using keras, this can make it follow scikit-learn's API.
- possibly slightly less flexible than TensorFlow.
The MLP base class¶
Note: I've removed some code and most documentation here for simplicity.
class MLPBaseEstimator(BaseEstimator, metaclass=ABCMeta):
def _preprocess_targets(self, y):
# Subclasses can override this to store information about the targets.
return y
def fit(self, X, y, monitor=None):
...
y = self._preprocess_targets(y)
self.graph_ = Graph()
with self.graph_.as_default():
# Define the model (We'll get to this in a minute).
self._init_model()
# Initialize weights.
self._session = tf.Session()
self._session.run(tf.initialize_all_variables())
...
# Minibatch training.
for epoch in range(self.n_epochs):
random_state.shuffle(indices)
for start_idx in range(0, n_examples, batch_size):
# Make the dictionary assigning a minibatch of training
# examples (pair of arrays) to TensorFlow placeholders.
batch_ind = indices[start_idx:start_idx + batch_size]
feed_dict = self._make_feed_dict(X[batch_ind],
y[batch_ind])
# Compute objective function and gradients and update weights.
obj_val, _ = self._session.run(
[self._obj_func, self._train_step],
feed_dict=feed_dict)
...
return self
Abstract methods to be implemented by the regressor and classifier classes¶
...
@abstractmethod
def _init_model_output(self, t):
pass
@abstractmethod
def _init_model_objective_fn(self, t):
pass
@abstractmethod
def predict(self, X):
pass
The core MLP modeling code¶
...
def _init_model(self):
# A placeholder variable to control dropout for training vs. prediction.
self._dropout = \
tf.placeholder(dtype=np.float32, shape=(), name="dropout")
# Input layers.
if self.is_sparse_:
...
else:
self._input_values = \
tf.placeholder(np.float32, [None, self.input_layer_sz_],
"input_values")
t = self._input_values
# Hidden layers (self.hidden_units is a list of ints for HL sizes)
for i, layer_sz in enumerate(self.hidden_units):
if self.is_sparse_ ...:
...
else:
t = tf.nn.dropout(t, keep_prob=self._dropout)
t = _affine(t, layer_sz, scope='layer_%d' % i)
t = t if self.activation is None else self.activation(t)
# The output layer and objective function depend on the model.
t = self._init_model_output(t)
self._init_model_objective_fn(t)
# Set the training algorithm, which is currently not configurable.
self._train_step = tf.train.AdamOptimizer().minimize(self._obj_func)
...
In [25]:
# You can't pickle some TensorFlow objects, at least as of version 0.10.0rc0.
import pickle
try:
g = tf.Graph()
pickle.dumps(g)
except TypeError:
print("You can't pickle a tf.Graph")
try:
with tf.Session() as sess:
pickle.dumps(sess)
except TypeError:
print("You can't pickle a tf.Session.")
You can't pickle a tf.Graph You can't pickle a tf.Session.
Methods to support pickling¶
- TensorFlow has a
Saverclass that writes models and parameters to disk. - One can override
__getstate__, the methodpickleuses to get an instance's data. - A not-so-elegent workaround to support TF pickling:
- write the model to disk
- read it in as bytes
- pickle
- unpickling is the reverse
...
# Used when saving:
def __getstate__(self):
# Write out the model.
...
if getattr(self, '_fitted', False):
tempfile = NamedTemporaryFile(delete=False)
tempfile.close()
try:
# Serialize the model and read it so it can be pickled.
self._saver.save(self._session, tempfile.name)
with open(tempfile.name, 'rb') as f:
saved_model = f.read()
finally:
os.unlink(tempfile.name)
...
# Note: don't include the graph since it can be recreated.
state = dict(
activation=self.activation,
batch_size=self.batch_size,
...
)
# Add fitted attributes if the model has been fit.
if getattr(self, '_fitted', False):
state['_fitted'] = True
state['input_layer_sz_'] = self.input_layer_sz_
state['is_sparse_'] = self.is_sparse_
state['_saved_model'] = saved_model
...
# Return what can and should be pickled.
return state
# Used when loading:
def __setstate__(self, state):
# Set hyperparameters, which pickled in the usual way.
for k, v in state.items():
if k in ['saved_model']:
continue
self.__dict__[k] = v
# Reinitialize a Graph and Session, and restore the saved values.
...
if state['_saved_model'] is not None:
tempfile = NamedTemporaryFile(delete=False)
tempfile.close()
try:
# Write out the serialized model that can be restored by TF.
with open(tempfile.name, 'wb') as f:
f.write(state['_saved_model'])
self.graph_ = Graph()
with self.graph_.as_default():
self._init_model()
self._session = tf.Session()
self._saver.restore(self._session, tempfile.name)
finally:
os.unlink(tempfile.name)
Classifier subclass¶
class MLPClassifier(MLPBaseEstimator, ClassifierMixin):
def __init__(self, hidden_units=(256,), batch_size=64, n_epochs=5,
dropout=None, activation=nn.relu, init_scale=0.1,
random_state=None):
self.hidden_units = hidden_units
self.batch_size = batch_size
...
def _init_model_output(self, t):
# Determine the output layer size.
if self.multilabel_:
...
elif self.n_classes_ > 2:
...
else:
# Binary classification
output_size = 1
# Add the final affine transformation.
if self.is_sparse_ ...:
...
else:
t = tf.nn.dropout(t, keep_prob=self._dropout)
t = _affine(t, output_size, scope='output_layer')
# Add the output layer activation function.
if self.multilabel_:
...
elif self.n_classes_ > 2:
...
else:
# Binary classification
self.input_targets_ = tf.placeholder(tf.int64, [None], "targets")
t = tf.reshape(t, [-1]) # Convert to 1d tensor.
self.output_layer_ = tf.nn.sigmoid(t)
return t
def _init_model_objective_fn(self, t):
if self.multilabel_:
...
elif self.n_classes_ > 2:
...
else:
# Binary classification
cross_entropy = tf.nn.sigmoid_cross_entropy_with_logits(
t, tf.cast(self.input_targets_, np.float32))
self._obj_func = tf.reduce_mean(cross_entropy)
def _preprocess_targets(self, y):
# Store a mapping between class label (e.g., strings) and indices.
...
def predict_proba(self, X):
...
def predict(self, X):
...
...
3. Simple distributed computing¶
motivation: ML methods have lots of hyperparameters.
- deep learning: numbers and sizes of hidden layers, etc.
- probabilistic graphical models: parameters for priors
- tree ensembles: depth, learning rate, samples per split, etc.
grid search
- try a lots of settings, pick the one leading to the best performance on held-out development data
- often combined with $k$-fold cross-validation (e.g., scikit-learn's
GridSearchCV) - crude but embarassingly parallel
Alternative hyperparameter search methods exist.
- randomized search, HyperOpt, BayesOpt, etc.
- also parallelizable to some extent
In [1]:
"""
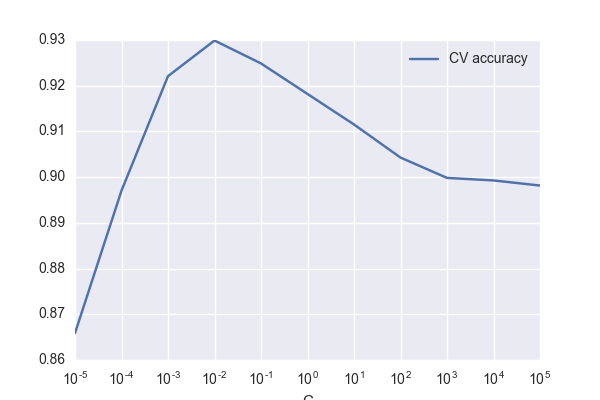
A realllllly simple illustration of how hyperparameter tuning matters.
Tuning the regularization for logistic regression on the "digits"
dataset in scikit-learn leads to a 14% error reduction in k-fold CV.
"""
import pandas as pd
from sklearn.datasets import load_digits
from sklearn.linear_model import LogisticRegression
from sklearn.grid_search import GridSearchCV
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import seaborn
digits = load_digits()
est = LogisticRegression(random_state=1234)
gs = GridSearchCV(est, param_grid={"C": [10.0 ** x for x in range(-5, 6)]})
gs.fit(digits.data, digits.target)
df = pd.DataFrame({'CV accuracy': [x.mean_validation_score
for x in gs.grid_scores_],
'C': [x.parameters['C'] for x in gs.grid_scores_]})
df.plot(x='C', y='CV accuracy', logx=True, figsize=(6, 4))
plt.savefig('hyperparameter_search.png')
default_score = gs.grid_scores_[5].mean_validation_score
print("error reduction (k-fold CV): {0:.0f}%"
.format(100 * (gs.best_score_ - default_score) / (1 - default_score)))
error reduction (k-fold CV): 14%
joblib¶
- a Python package for running embarassingly parallel tasks
- contributors: @GaelVaroquaux, @ogrisel, et al.
- originally focused on single machine parallelization
- backends for multiprocessing, multithreading
A simple joblib example¶
- Select a backend (or use the default multiprocessing backend).
- Instantiate a
Parallelcallable. - Call it on a bunch of
delayedfunction calls and their arguments.
In [26]:
from joblib.parallel import delayed, parallel_backend, Parallel
def foo(x):
return 10 * x ** 2
with parallel_backend("multiprocessing"):
parallel = Parallel()
numbers = range(10)
print("map a function f(x)=10*x**2 over a list of numbers:\n",
parallel(delayed(foo)(x) for x in numbers))
map a function f(x)=10*x**2 over a list of numbers: [0, 10, 40, 90, 160, 250, 360, 490, 640, 810]
joblib in scikit-learn¶
- scikit-learn uses joblib to parallelize hyperparameter evaluation in
GridSearchCV - If you tell joblib to use a custom backend, scikit-learn will use that for grid search.
joblib custom backends¶
- version 0.10.0 supports custom parallel backends
- example: dask distributed backend
implementing a custom backend¶
- a class representing a pending result
- similar to a
concurrent.Future getmethod to block and retrieve the resultresultattribute to cache the result
- similar to a
- a
joblib._parallel_backends.ParallelBackendBasesubclasseffective_n_jobsmethod that determines how many jobs can run in parallel- e.g., can be used to cap the number of cores used across a cluster
apply_asyncmethod- queues up computation of a function call
- returns a result instance (see above)
Civis's computation infrastructure¶
- Civis has an autoscaling EC2- and Docker-based system for running jobs.
- alternatives: kubernetes, mesos
- Jobs are independent (currently, at least).
- We have a Python client with a
concurrent.futures.Executorinterface.- This allows us to submit docker run commands to remote workers.
- We'll use a joblib backend to facilitate getting functions and data to the workers.
Outline of our backend implementation¶
class _CivisBackend(ParallelBackendBase):
def __init__(self, ...):
...
# Initialize an executor for making and running Docker containers.
self.executor = ContainerPoolExecutor(**executor_kwargs)
...
def effective_n_jobs(self, n_jobs):
# e.g., set a hard limit on the number of jobs.
...
def apply_async(self, func, callback=None):
...
with TemporaryDirectory() as tempdir:
# Serialize func to a temporary file and upload it to Civis.
...
# Make a command for the remote Docker container that will run
# a script that will download the serialized function `func`,
# run it, and store the result.
cmd = (...
"python {runner_script} {func_file_id}"
...)
...
# Submit the command to be executed in a remote Docker container.
future = self.executor.submit(cmd)
...
# Wait for the job to finish.
result = _CivisFutureResult(future, callback)
return result
class _CivisFutureResult:
def __init__(self, future, callback):
...
def get(self):
if self.result is None:
...
# Wait for the script to complete.
self._future.result()
...
# Download and deserialize the result.
with TemporaryDirectory() as tempdir:
temppath = os.path.join(tempdir, "civis_joblib_backend_result")
# Download the serialized result.
...
self.result = joblib.load(temppath)
...
return self.result
Does it work?¶
- Evaluate the MLP vs. logistic regression in terms of model accuracy.
- Evaluate grid searching on my laptop versus distributed in the #cloud in terms of wall time.
A basic evaluation for text classification¶
- Pang & Lee (ACL 2004) Movie Reviews dataset
- 2,000 movie reviews with positive/negative (0/1) sentiment polarity labels
- features
- character $n$-grams from sklearn's
CountVectorizer
- character $n$-grams from sklearn's
- Evaluation setup:
- randomly split 75% for training, 25% for testing
- hyperparameter grid search on the training set
- evaluate with the ROC AUC score of testing set predictions
In [1]:
import nltk
from nltk.corpus import movie_reviews
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
from sklearn.cross_validation import ShuffleSplit
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.grid_search import GridSearchCV
from sklearn.metrics import get_scorer
from muffnn import MLPClassifier
In [2]:
# Load the movie reviews data
# (polarity dataset v2.0 here: http://www.cs.cornell.edu/people/pabo/movie-review-data/).
nltk.download('movie_reviews')
X = np.array([movie_reviews.raw(i) for i in movie_reviews.fileids()])
y = np.array([1 if x.split('/')[0] == 'pos' else 0
for x in movie_reviews.fileids()])
splits = ShuffleSplit(X.shape[0], n_iter=1, test_size=0.25, random_state=1234)
train_ind, test_ind = [x for x in splits][0]
[nltk_data] Downloading package movie_reviews to [nltk_data] /Users/civisemployee/nltk_data... [nltk_data] Package movie_reviews is already up-to-date!
In [3]:
# character n-grams and a Multilayer Perceptron classifier
ct_vect = CountVectorizer(analyzer='char', ngram_range=(2, 5),
max_features=50000)
mlp = MLPClassifier(n_epochs=10, random_state=42)
pipeline = Pipeline(steps=[('char_ngram', ct_vect),
('mlp', mlp)])
param_grid = {
'mlp__hidden_units': [(512,), (256,), (256, 128, 64)],
'mlp__dropout': [None, 0.5]
}
gs_mlp = GridSearchCV(pipeline, param_grid=param_grid,
n_jobs=4, scoring='roc_auc')
In [4]:
# baseline: character n-grams and a logistic regression classifier
ct_vect = CountVectorizer(analyzer='char', ngram_range=(2, 5),
max_features=50000)
lr = LogisticRegression(random_state=42)
pipeline = Pipeline(steps=[('char_ngram', ct_vect),
('lr', lr)])
param_grid = {
'lr__C': [0.001, 0.01, 0.1, 1.0, 10.0, 100.0]
}
gs_lr = GridSearchCV(pipeline, param_grid=param_grid,
n_jobs=4, scoring='roc_auc')
In [ ]:
%time gs_lr.fit(X[train_ind], y[train_ind])
%time gs_mlp.fit(X[train_ind], y[train_ind])
CPU times: user 27.5 s, sys: 876 ms, total: 28.3 s
Wall time: 4min
CPU times: user 9min 1s, sys: 4min 54s, total: 13min 55s
Wall time: 51min 32sIn [ ]:
roc_auc_scorer = get_scorer('roc_auc')
print("Logistic Regression ROC AUC: {:.4f}".format(
roc_auc_scorer(gs_lr, X[test_ind], y[test_ind])))
print("MLP ROC AUC: {:.4f}".format(
roc_auc_scorer(gs_mlp, X[test_ind], y[test_ind])))
Logistic Regression ROC AUC: 0.9119
MLP ROC AUC: 0.9271With distributed grid search¶
factory = make_backend_factory(
required_resources={"cpu": 2048, "memory": 4096}, ...)
register_parallel_backend('civis', factory)
with parallel_backend('civis'):
gs_mlp.fit(X[train_ind], y[train_ind])
Wall time:
- laptop: 51min 32s
- distributed: 16min 23s
Notes: both include refitting on all the data locally after grid search.
Parting thoughts (speculation)¶
- With deep learning, speeding up hyperparameter search is important.
- Distributed grid search > distributed model fitting
- (probably, currently, for many applications)
- When combining open-source tools,
whole > np.sum(parts)
Thanks¶
- Matt Becker, Bill Lattner, Derrick Higgins (for code review)
- Walt Askew (for code contributions)